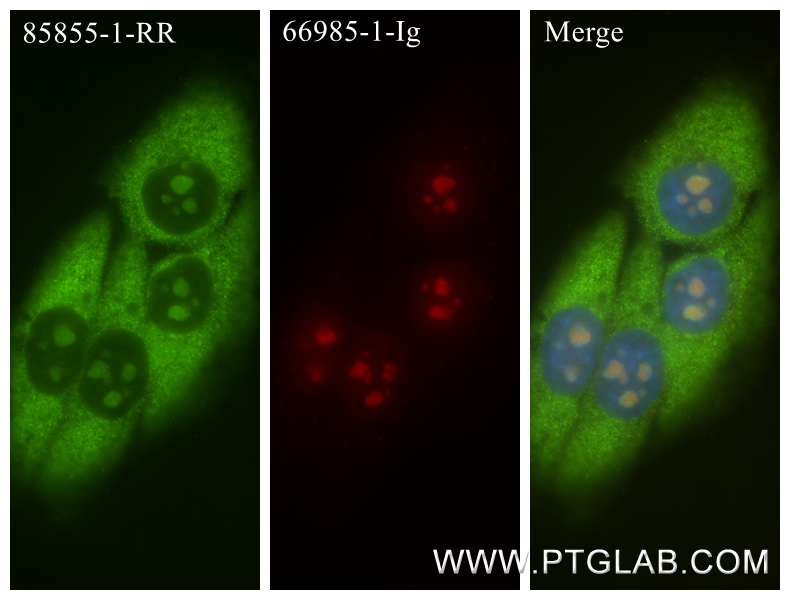
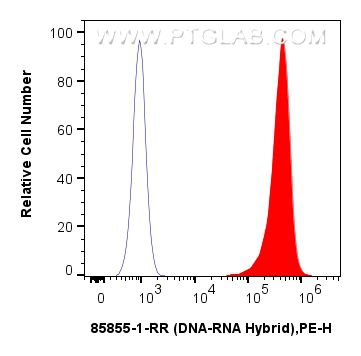
验证数据展示
经过测试的应用
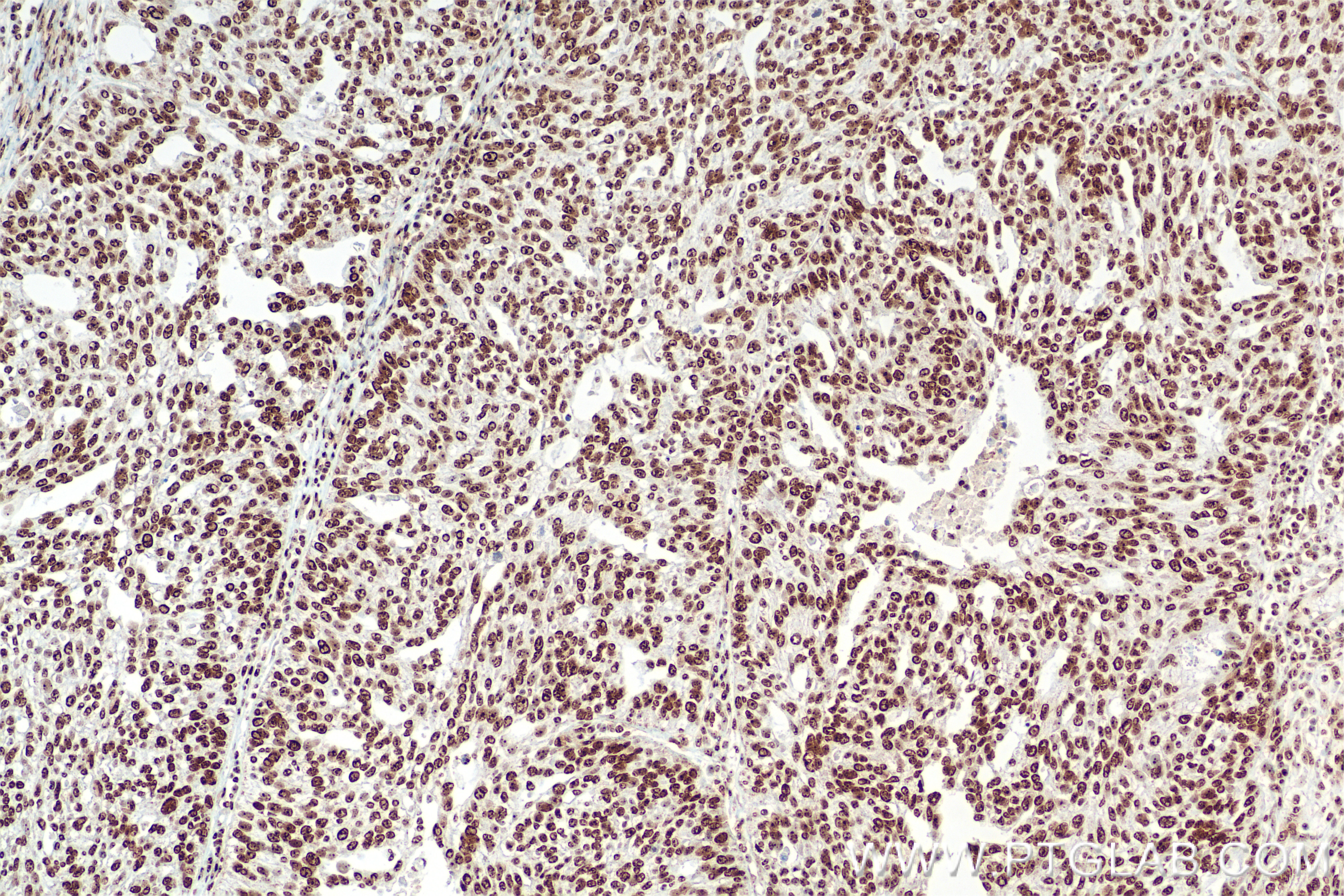
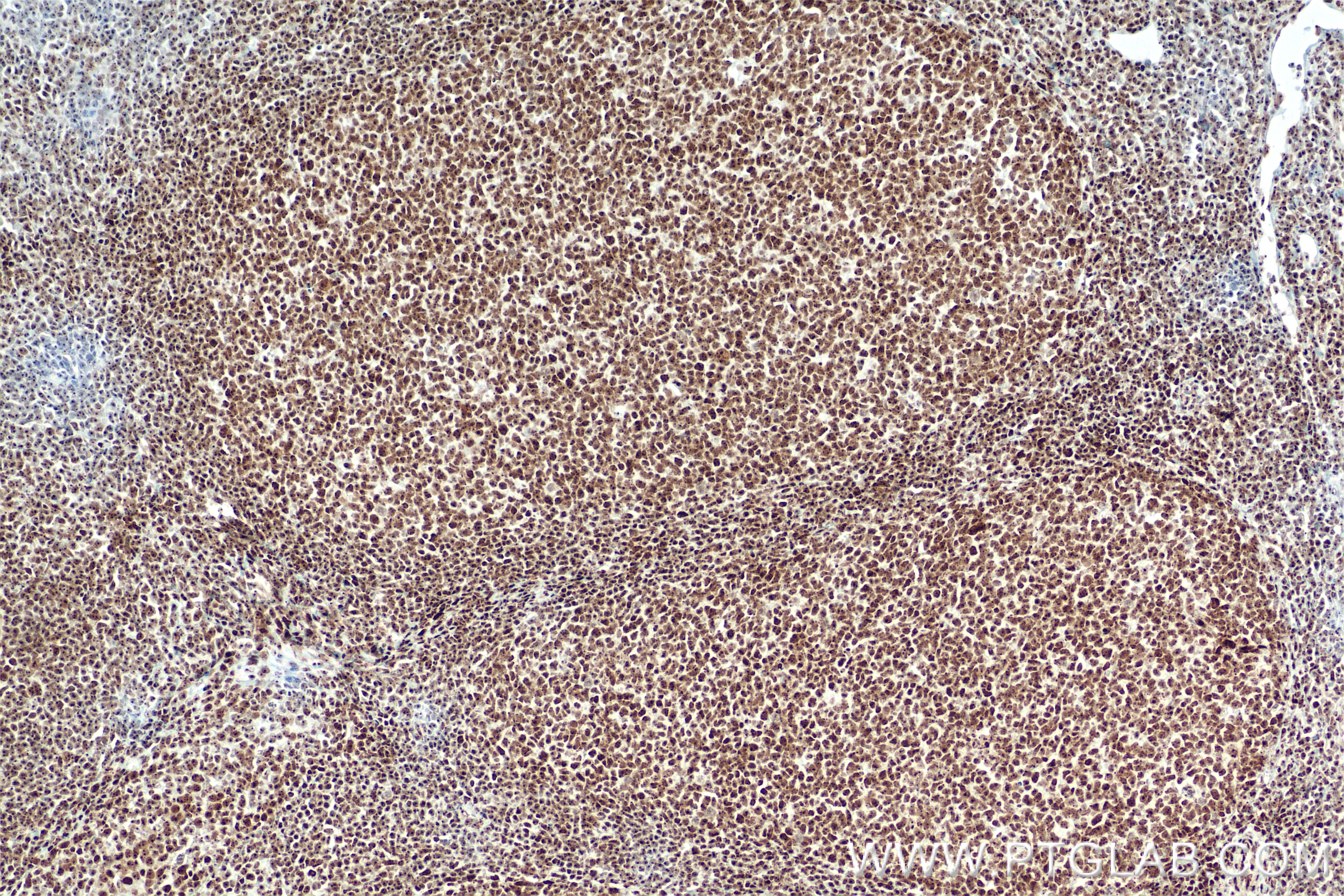
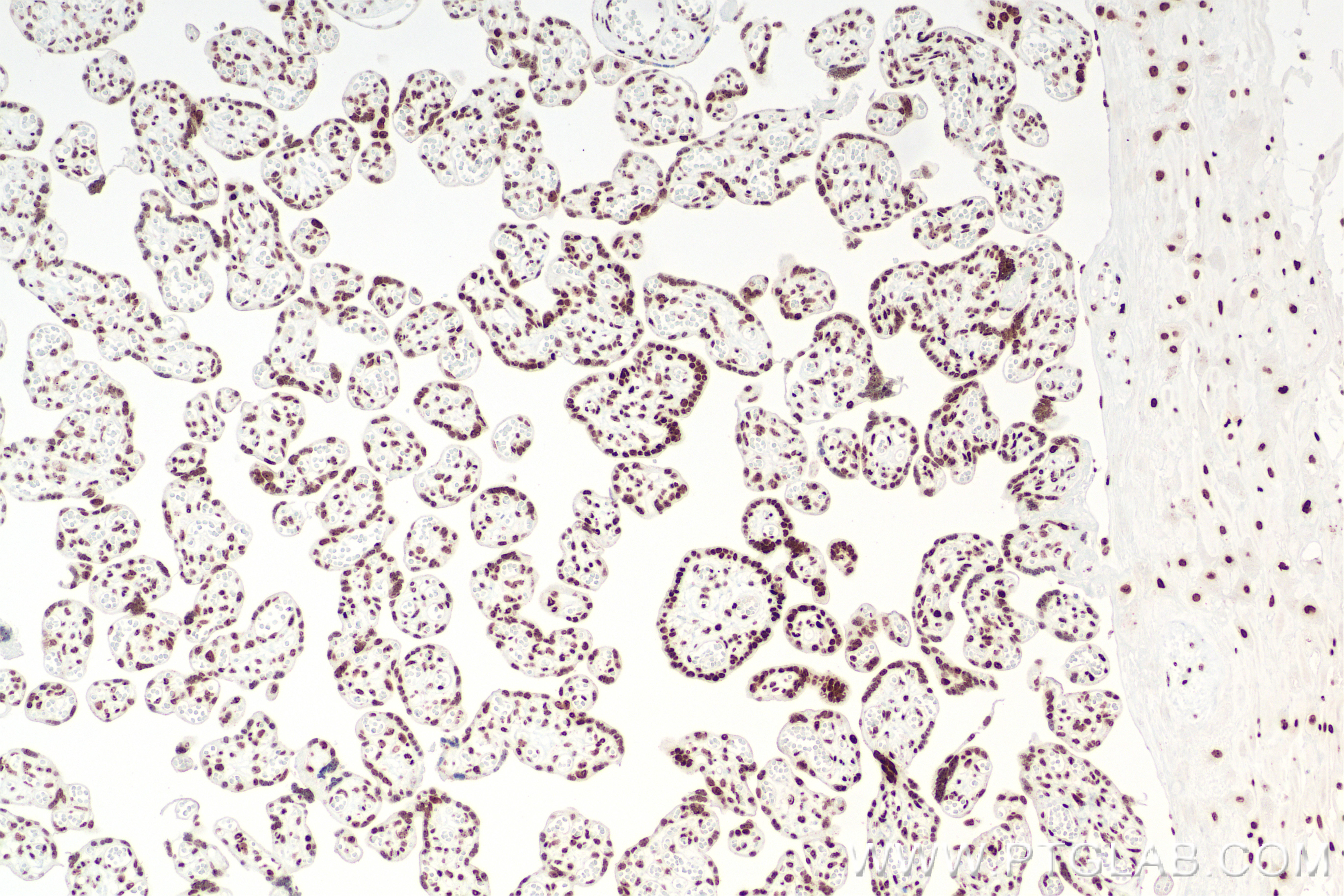
| Positive IHC detected in | human placenta tissue, human ovary tumor tissue, human tonsillitis tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
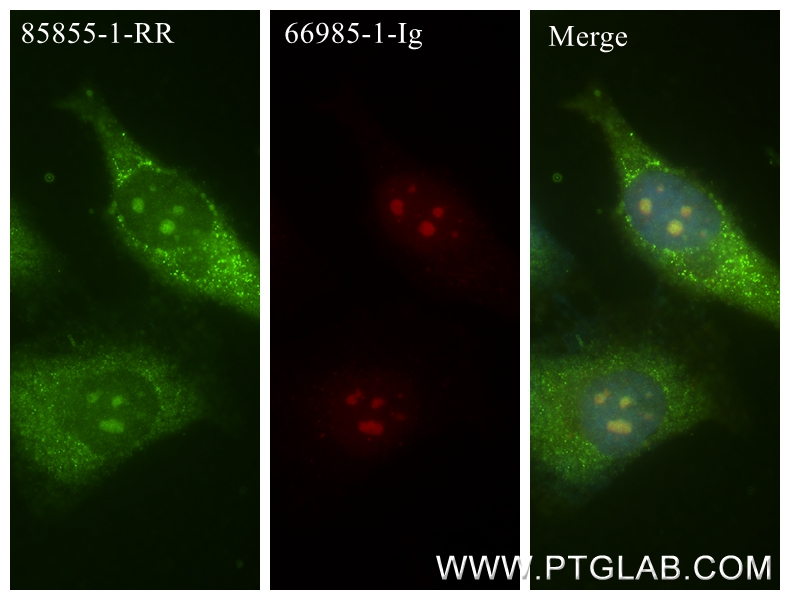
| Positive IF/ICC detected in | HepG2 cells, HeLa cells |
| Positive FC (Intra) detected in | HEK-293T cells |
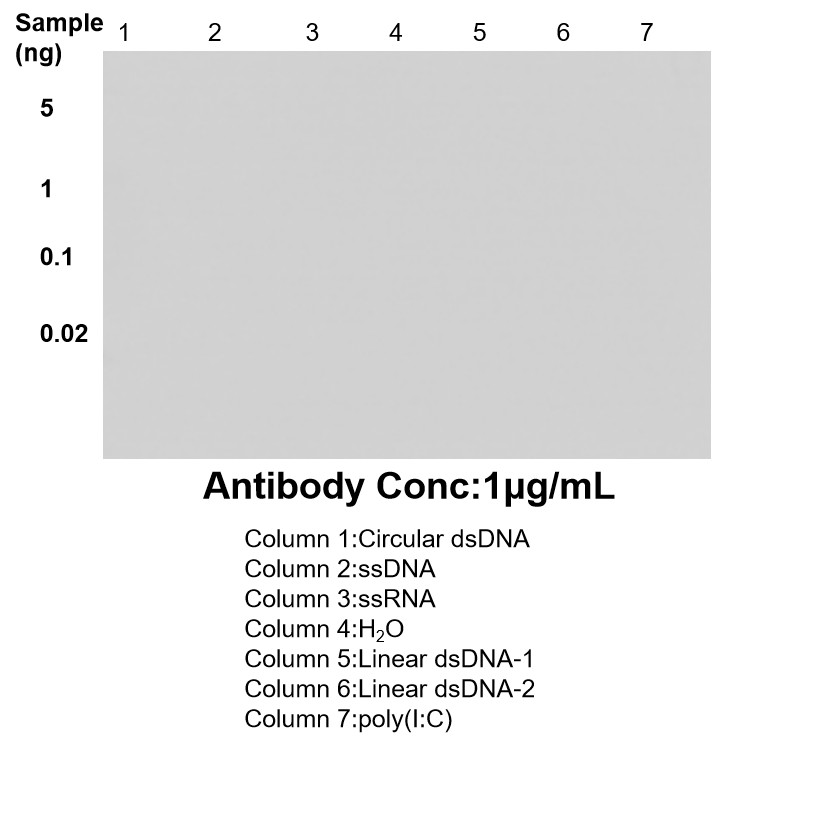
| Positive Dot blot detected in | / / |
推荐稀释比
| 应用 | 推荐稀释比 |
|---|---|
| Immunohistochemistry (IHC) | IHC : 1:400-1:1600 |
| Immunofluorescence (IF)/ICC | IF/ICC : 1:125-1:500 |
| Flow Cytometry (FC) (INTRA) | FC (INTRA) : 0.2 ug per 10^6 cells in a 100 µl suspension |
| DOT BLOT | DOT BLOT : 1:10-1:100 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
产品信息
85855-1-RR targets DNA-RNA Hybrid in IHC, IF/ICC, FC (Intra), Dot blot, ELISA applications and shows reactivity with n/a samples.
| 经测试应用 | IHC, IF/ICC, FC (Intra), Dot blot, ELISA Application Description |
| 经测试反应性 | n/a |
| 免疫原 |
unknow 种属同源性预测 |
| 宿主/亚型 | Rabbit / IgG |
| 抗体类别 | Recombinant |
| 产品类型 | Antibody |
| 全称 | DNA-RNA Hybrid |
| 别名 | |
| 基因名称 | |
| Gene ID (NCBI) | |
| RRID | AB_3744425 |
| 偶联类型 | Unconjugated |
| 形式 | Liquid |
| 纯化方式 | Protein A purification |
| 储存缓冲液 | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| 储存条件 | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. |
背景介绍
DNA-RNA Hybrid antibodies are primarily used to study R-loops. Theoretically, they do not bind to single-stranded DNA, double-stranded DNA, or RNA. R-loops are DNA-RNA hybrid structures formed during transcription and are closely related to genome stability, DNA replication, DNA repair, and cancer development.
实验方案
| Product Specific Protocols | |
|---|---|
| IF protocol for DNA-RNA Hybrid antibody 85855-1-RR | Download protocol |
| IHC protocol for DNA-RNA Hybrid antibody 85855-1-RR | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |