验证数据展示
经过测试的应用
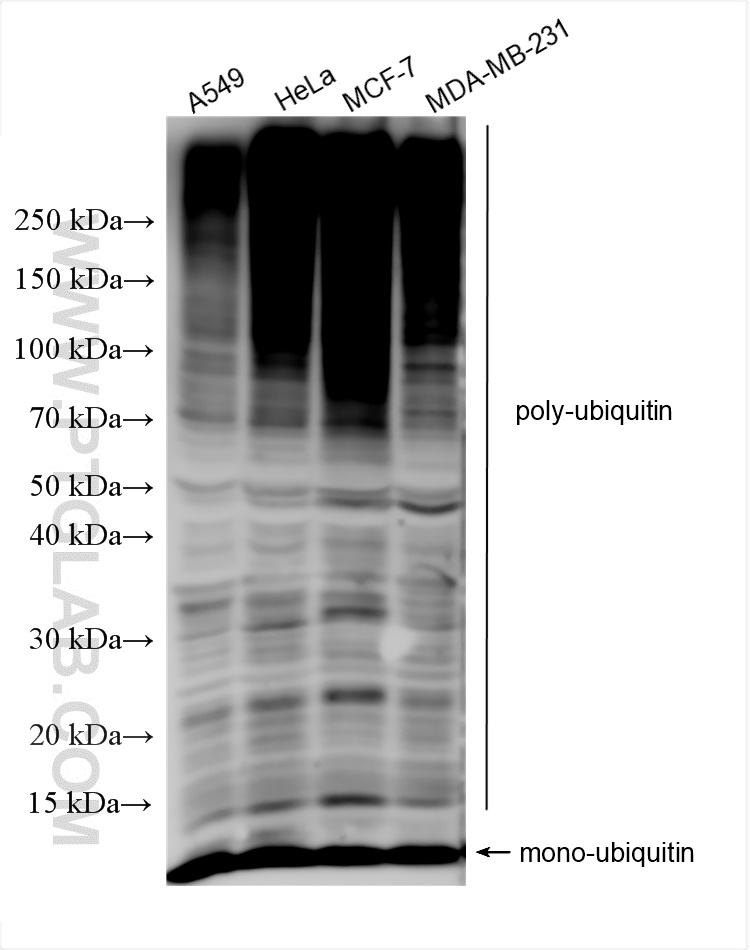
| Positive WB detected in | A549 cells, HeLa cells, MCF-7 cells, MDA-MB-231 cells |
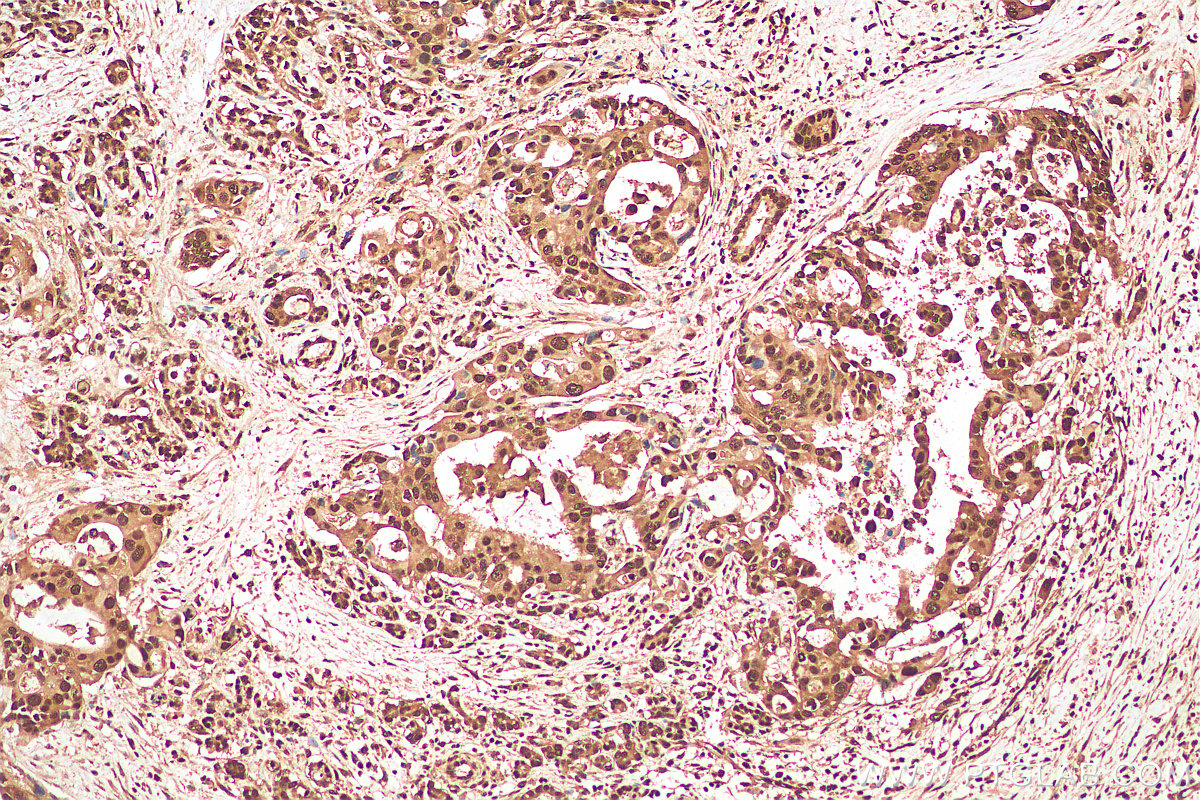
| Positive IHC detected in | human pancreas cancer tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
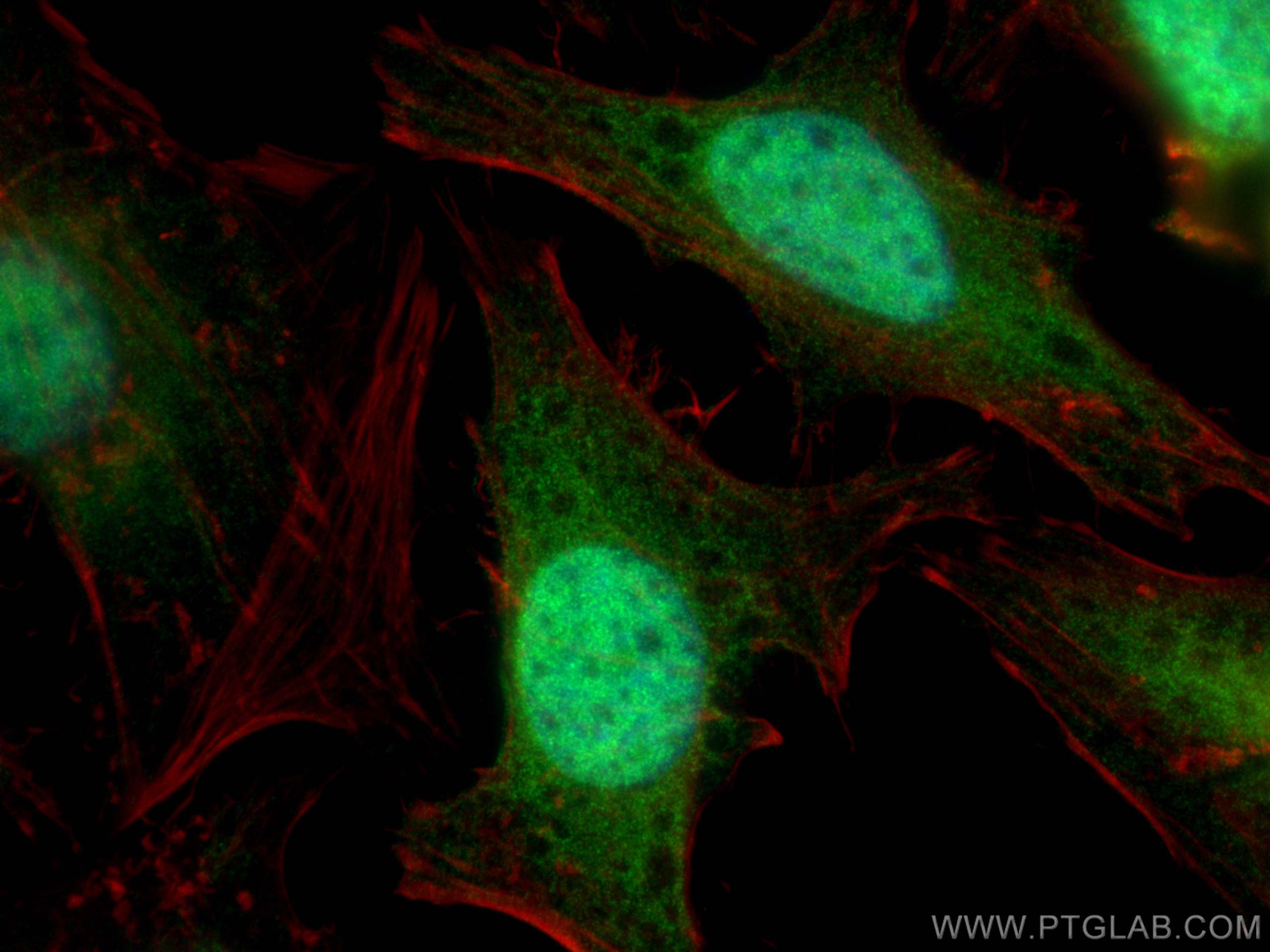
| Positive IF/ICC detected in | HeLa cells |
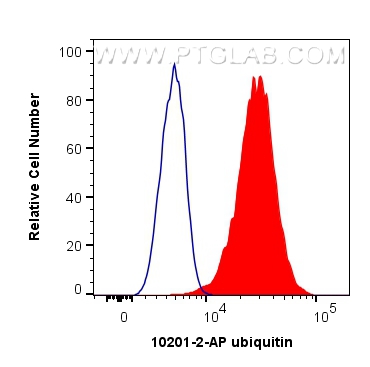
| Positive FC (Intra) detected in | HeLa cells |
推荐稀释比
| 应用 | 推荐稀释比 |
|---|---|
| Western Blot (WB) | WB : 1:1000-1:8000 |
| Immunohistochemistry (IHC) | IHC : 1:50-1:500 |
| Immunofluorescence (IF)/ICC | IF/ICC : 1:200-1:800 |
| Flow Cytometry (FC) (INTRA) | FC (INTRA) : 0.40 ug per 10^6 cells in a 100 µl suspension |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
产品信息
10201-2-AP targets ubiquitin in WB, IHC, IF/ICC, FC (Intra), CoIP, ChIP, ELISA applications and shows reactivity with human, mouse, rat samples.
| 经测试应用 | WB, IHC, IF/ICC, FC (Intra), ELISA Application Description |
| 文献引用应用 | WB, IHC, IF, CoIP, ChIP |
| 经测试反应性 | human, mouse, rat |
| 文献引用反应性 | human, mouse, rat, canine, monkey, chicken, arabidopsis, trypanosoma cruzi, c.elegans |
| 免疫原 |
CatNo: Ag0260 Product name: Recombinant human ubiquitin protein Source: e coli.-derived, PGEX-4T Tag: GST Domain: 104-229 aa of BC000379 Sequence: AKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYNIQKESTLHLVLRLRGGMQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYNIQKESTLHLVLRLRGGC 种属同源性预测 |
| 宿主/亚型 | Rabbit / IgG |
| 抗体类别 | Polyclonal |
| 产品类型 | Antibody |
| 全称 | ubiquitin B |
| 别名 | UBB, ubiquitin B, Polyubiquitin B, Polyubiquitin-B |
| GenBank蛋白编号 | BC000379 |
| 基因名称 | ubiquitin |
| Gene ID (NCBI) | 7314 |
| RRID | AB_671515 |
| 偶联类型 | Unconjugated |
| 形式 | Liquid |
| 纯化方式 | Antigen affinity purification |
| UNIPROT ID | P0CG47 |
| 储存缓冲液 | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| 储存条件 | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. |
背景介绍
Ubiquitin B (UBB) is a member of ubiquitin family, one of the most conserved proteins known. Ubiquitin B is required for ATP-dependent, non-lysosomal intracellular protein degradation of abnormal proteins and normal proteins with a rapid turnover. Ubiquitin B is covalently bound to proteins to be degraded, and presumably labels these proteins for degradation. Ubiquitin also binds to histone H2A in actively transcribed regions but does not cause histone H2A degradation, suggesting that ubiquitin is also involved in regulation of gene expression.When polyubiquitin is free (unanchored-polyubiquitin), it also has distinct roles, such as in activation of protein kinases, and in signaling. This gene consists of three direct repeats of the ubiquitin coding sequence with no spacer sequence. Consequently, the protein is expressed as a polyubiquitin precursor with a final amino acid after the last repeat. Aberrant form of this protein has been noticed in patients with Alzheimer's and Down syndrome. Interestingly ubiquitin also becomes covalently bonded to many types of pathological inclusions which appear to be resistant to normal degradation.
实验方案
| Product Specific Protocols | |
|---|---|
| FC protocol for ubiquitin antibody 10201-2-AP | Download protocol |
| IF protocol for ubiquitin antibody 10201-2-AP | Download protocol |
| IHC protocol for ubiquitin antibody 10201-2-AP | Download protocol |
| WB protocol for ubiquitin antibody 10201-2-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
发表文章
| Species | Application | Title |
|---|---|---|
Nature Proteasome inhibition for treatment of leishmaniasis, Chagas disease and sleeping sickness. | ||
Cell Res NudCL2 is an autophagy receptor that mediates selective autophagic degradation of CP110 at mother centrioles to promote ciliogenesis. | ||
Nat Commun UV-B irradiation-activated E3 ligase GmILPA1 modulates gibberellin catabolism to increase plant height in soybean | ||
Nat Commun Stabilization of Pin1 by USP34 promotes Ubc9 isomerization and protein sumoylation in glioma stem cells | ||
Nat Commun Smooth muscle NF90 deficiency ameliorates diabetic atherosclerotic calcification in male mice via FBXW7-AGER1-AGEs axis |